pLM4CPPs is a deep learning architecture designed for predicting cell-penetrating peptides (CPPs). At its core, pLM4CPPs utilizes advanced pretrained protein language models (pLMs) trained on extensive protein sequence data. These models capture intricate sequence relationships and functional motifs critical for CPP activity, enhancing accuracy and reliability in classification. Key to pLM4CPPs is its integration of Convolutional Neural Networks (CNNs) for hierarchical feature extraction from peptide sequences, achieving superior performance metrics such as accuracy, Matthews Correlation Coefficient (MCC), and sensitivity. Multiple peptide embeddings from sources like BEPLER, CPCProt, SeqVec, ESM variants (ESM, ESM-2, ESM-1b, ESM-1v), ProtT5-XL UniRef50, and ProtT5-XL BFD are evaluated to optimize performance across diverse datasets. pLM4CPPs integrates predictions from multiple models to provide a consensus decision on CPP classification, ensuring robust results and reliability. This platform is the implementation of the paper: Kumar, N.; Du, Z.; Li, Y. pLM4CPPs: Protein Language Model-Based Predictor for Cell Penetrating Peptides, J. Chem. Inf. Model. 2024 (Submitted).
Quick Output Version:
Large-scale Output Version:
Usage of the Web Server:
Quick Output Version:
Select a model, input peptide sequences, and click "Run" for quick predictions.Notice: Support multiple sequences input (e.g., "VPP,IPP,CCL,AGR").
Large-scale Output Version:
Upload files (xls, xlsx, txt, fasta) and click "Run" for batch predictions.Notice: File preparation guidelines available at the repository.
Model Performance on Test Dataset:
| Model | ACC | BACC | Sn | Sp | MCC |
|---|---|---|---|---|---|
| pLM4CPPs (ESM-1280) | 0.929 | 0.893 | 0.820 | 0.966 | 0.808 |
| pLM4CPPs (ESM-640) | 0.923 | 0.880 | 0.791 | 0.968 | 0.792 |
| pLM4CPPs (ESM-480) | 0.931 | 0.907 | 0.860 | 0.955 | 0.816 |
| pLM4CPPs (ESM-320) | 0.923 | 0.892 | 0.831 | 0.955 | 0.795 |
| pLM4CPPs (ProtT5-XL BFD) | 0.921 | 0.891 | 0.831 | 0.951 | 0.789 |
| pLM4CPPs (SeqVec) | 0.932 | 0.901 | 0.838 | 0.965 | 0.819 |
Schematic framework of pLM4CPPs:
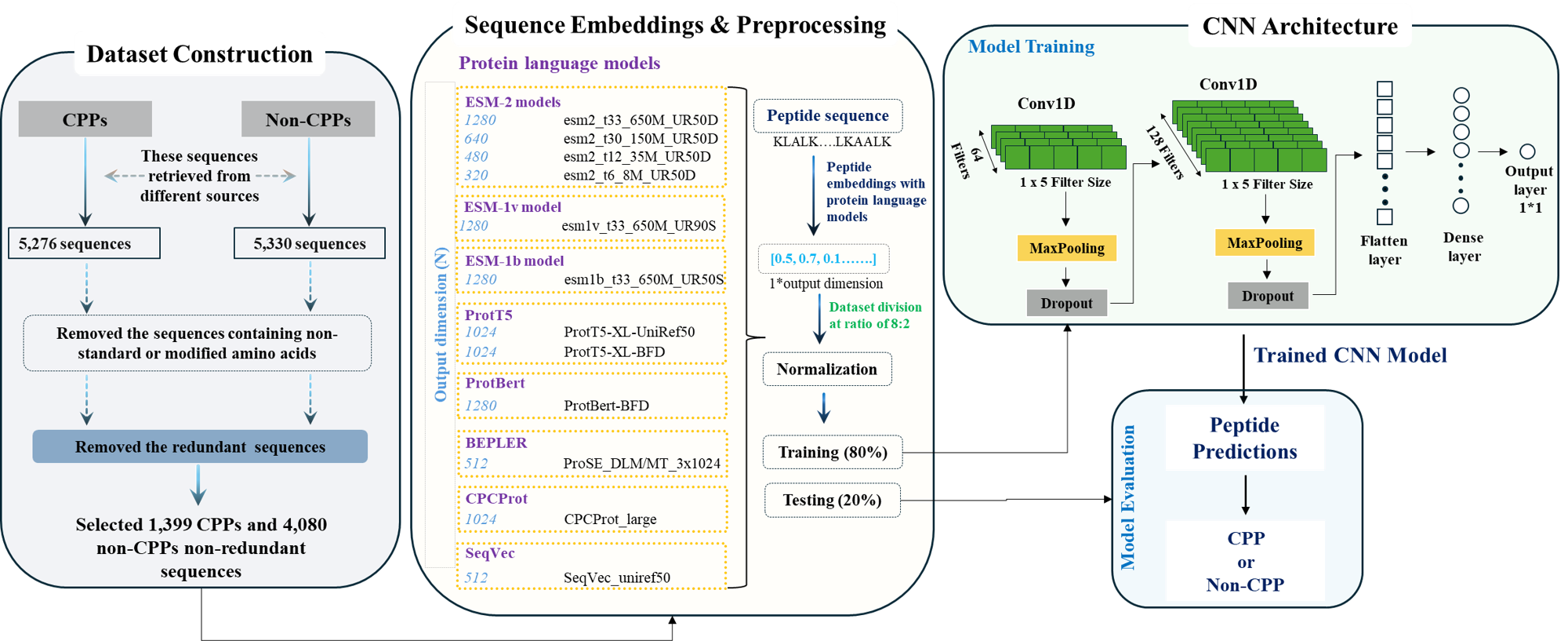
Contact and Support
Nandan Kumar (nandan@ksu.edu) Zhenjiao Du (zhenjiao@ksu.edu) Yonghui Li (yonghui@ksu.edu)